Invited Symposium: Intracellular Traffic of Organelles
| INABIS '98 Home Page | Your Session | Symposia & Poster Sessions | Plenary Sessions | Exhibitors' Foyer | Personal Itinerary | New Search |
Results
SDS-Resistant SNARE Complexes Are Present in Drosophila.
SNARE proteins interact with each other to promote membrane fusion, forming post-fusion SDS-resistant complexes which when dissociated through the catalytic action of NSF and SNAP (18, 19), may be recycled back into the transport pathway. In attempts to use Drosophila as a model system for studying membrane transport and exocytosis, Drosophila NSF, SNAP and SNARE proteins have been identified. Advances have been made in determining the function of dSNAREs, as mutations in dsyntaxin and deactivation of dVAMP severely inhibit neurotransmission (42). However, the function of dNSF and dSNAP has not yet been determined.
To determine whether dNSF and dSNAP are capable of dissociating the 7S complex, we first began by testing their ability to dissociate mammalian 7S complex. Methods have been established to permit the formation and dissociation of 7S complexes formed from recombinant proteins in vitro (19). Such cross-species experiments are not unprecedented, specially in the transport system where yeast SNAP and NSF were able to substitute for mammalian SNAP in intra-Golgi transport assay (28). A fusion protein consisting of glutathione-S-transferase (GST) at the N-terminal coupled to the cytoplasmic domain of syntaxin (1-267) was used as the central protein to bind other recombinant proteins which were all his-tagged at the N-terminal. Upon incubation of GST-syntaxin bound to glutathione-conjugated beads with the cytoplasmic portion of VAMP-2 (1-94), and his-SNAP-25, SDS resistant complexes assembled spontaneously. Figure 1 shows that a portion of syntaxin protein is in an SDS resistant complex with SNAP-25 and VAMP-2 (lane4,N). In contrast, syntaxin is incapable of forming such a complex by itself (lane2,N). Boiled samples show GST-syntaxin in monomeric form (lanes 1,B and 3,B).
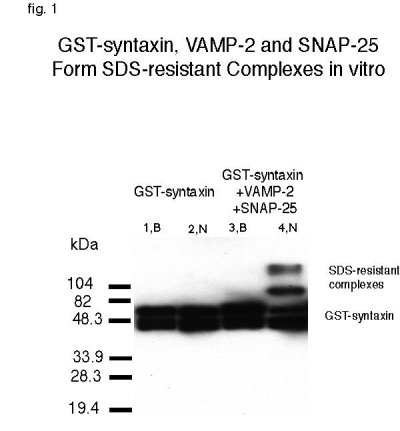 Fig. 1:GST-syntaxin, VAMP-2 and SNAP-25 form SDS-resistant complexes in vitro. 0.1 nmole of GST-syntaxin was incubated by itself or with 0.5 nmole of VAMP and SNAP-25. Samples were then placed in loading buffer containing 1% SDS and electrophoresed before (N) or after (B) boiling. Anti-syntaxin was used to detect SDS-resistant complexes in lane 4 and monomeric GST-syntaxin in all lanes.
Fig. 1:GST-syntaxin, VAMP-2 and SNAP-25 form SDS-resistant complexes in vitro. 0.1 nmole of GST-syntaxin was incubated by itself or with 0.5 nmole of VAMP and SNAP-25. Samples were then placed in loading buffer containing 1% SDS and electrophoresed before (N) or after (B) boiling. Anti-syntaxin was used to detect SDS-resistant complexes in lane 4 and monomeric GST-syntaxin in all lanes.
In order for dNSFs to be capable of uncoupling the mammalian 7S complex, it would require the adaptor protein, dSNAP, to bind to mammalian syntaxin in the complex and create a site for dNSF attachment (43). Recombinant his-dSNAP was purified and incubated with GST-syntaxin but was unable to bind even at concentrations fifty times higher that of syntaxin�s, despite 62% identity with mammalian SNAP (data not shown)(41). As positive control, mammalian his-SNAP binding to syntaxin was demonstrated (fig. 2b).
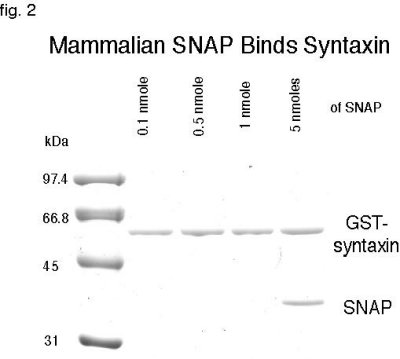 Fig. 2: Mammalian SNAP Binds to Syntaxin. 0.1 nmole of GST-syntaxin was incubated with different concentrations of SNAP. Coomassie staining demonstrates binding with 1 nmole of SNAP but the binding is much stronger at 5 nmoles.
Fig. 2: Mammalian SNAP Binds to Syntaxin. 0.1 nmole of GST-syntaxin was incubated with different concentrations of SNAP. Coomassie staining demonstrates binding with 1 nmole of SNAP but the binding is much stronger at 5 nmoles.
Since dSNAP could not interact with mammalian syntaxin, it led us to search for a source of 7S complex in Drosophila that is more likely serve as a receptor for dSNAP. First, we wanted to determine whether SNARE proteins behave biochemically similar to their mammalian counterpart in forming 7S complexes resistant to 1% SDS treatment. Fly lysates were prepared and fractionated according to material and methods (35). Electrophoretic separation of each fraction in SDS polyacrylamide gel resulted in detection of several bands by anti-dsyntaxin (8C3) antibody (fig. 3). Two major bands are present in the unboiled sample, one corresponding to the monomer syntaxin at 36 kDa (arrow) and another at 70 kDa (arrowhead) which represents the 7S complex, the sum of dsyntaxin, dSNAP-25 and dVAMP molecular weights (MW). Bands with MW higher than that of 70 kDa (*) may be multimers of the 7S complex. Boiling of the samples before electrophoresis, resulted in the dissociation of the complex and the appearance of a single syntaxin band at its monomeric molecular weight (fig. 3, lanes designated +). Future experiments will examine the ability of exogenously added dNSF and dSNAP to disassemble these SDS-resistant complexes endogenously present in lysates. We will also attempt to assemble recombinant Drosophila SNARE proteins into SDS-resistant complexes and study their dissociation in vitro.
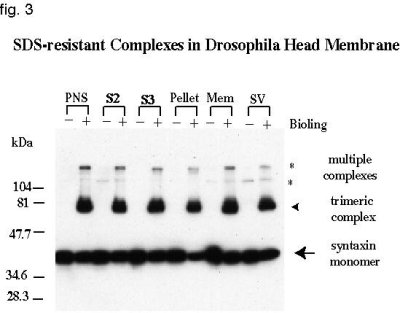 Fig. 3: SDS-resistant Complexes in Drosophila Head membrane. Anti-dsyntaxin (8C3) was used to detect the presence of SDS-resistant complexes in different fractions of fly lysate.
Fig. 3: SDS-resistant Complexes in Drosophila Head membrane. Anti-dsyntaxin (8C3) was used to detect the presence of SDS-resistant complexes in different fractions of fly lysate.
Nature of comatose mutation.
Concurrent with the in vitro studies, we are investigating the function of dNSF-1 in vivo, using a temperature-sensitive allele of dNSF-1 mutant line, comatose TP7. The severity of the TP7 allele was examined by placing the flies homozygous for the mutation in non-permissive temperatures of 38 oC. Within 2 minutes of heat-shock, the flies became paralyzed, a rate similar to other temperature-sensitive alleles of comatose (38). Also similar to other comatose alleles, it took 30 minutes for the flies to recover from paralysis after they were moved back to room temperature (37).
The nature of the TP7 mutation in dNSF-1 is not known; but if the function of dNSF-1 is to dissociate SNARE complexes, then one may expect to find less monomeric forms of SNARE proteins and an increased amount of SDS resistant complexes in comatose flies under non-permissive temperature of 38 oC. Comatose and wild-type Oregon R flies were heat shock treated for 4 minutes and immediately placed on dry ice where the heads were isolated and homogenized. Conventionally, to study 7S complexes, membrane homogenates are solubilized by non-ionic detergents such as Triton-X100 which promotes 7S complex assembly even after solubilization (44). To prevent formation of complexes during solubilization, we added 0.5% SDS to the post-nuclear supernatant (PNS) (20). SDS-resistant complexes formed before their exposure to SDS will remain in complex, but SDS prevent formation of new ones. 5mg samples of PNS were electrophoresed in each lane of the polyacrylamide gel and probed with anti-dsyntaxin. As controls, samples of non-heat-shock treated Oregon R and comatose flies were used. As expected, there was no difference between the heat shock treatment of Oregon R and control non-treated flies as detected with anti-dsyntaxin (fig.4, lanes 2 and 4). Surprisingly, however, there was also no discernible difference in formation of 7S complexes or the amount of monomeric form of syntaxin when comparing heat-shock treated and non-heat shock treated comatose flies (fig. 4, lanes 6 and 8).
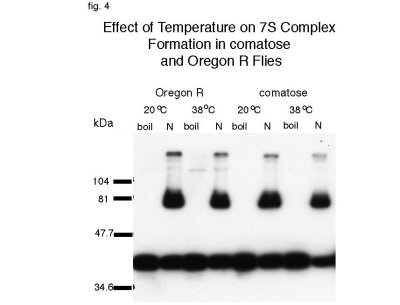 Fig. 4: Effect of Temperature on 7S Complex Formation in comatose and Oregon R Flies. Incubation of comatose flies at non-permissive temperature had no detectable effect on formation of 7S complexes as probed by anti-dsyntaxin (8C3)in vivo
Fig. 4: Effect of Temperature on 7S Complex Formation in comatose and Oregon R Flies. Incubation of comatose flies at non-permissive temperature had no detectable effect on formation of 7S complexes as probed by anti-dsyntaxin (8C3)in vivo
To precisely characterize the TP7 allele, we determined the sequence of TP7 dNSF-1 coding region. RT-PCR was performed using oligonucleotides flanking the dNSF-1 coding sequence as described in Material and Methods. Two independent PCR reactions produced bands of 2.2 kb, the same size as the band obtained by using dNSF-1 cDNA in plasmid DNA (control lane). Upon subcloning and examining their restriction patterns, the PCR products were sequenced. As predicted for a temperature-sensitive mutation, only a single point mutation was found. A change in amino acid number 397 from a proline to a serine residue appears to be responsible for the phenotype (fig. 5). Proline 397 is conserved throughout evolution and is located in the D1 domain, the region which has been demonstrated to be responsible for the ATPase activity in the mammalian NSF (3, 45).
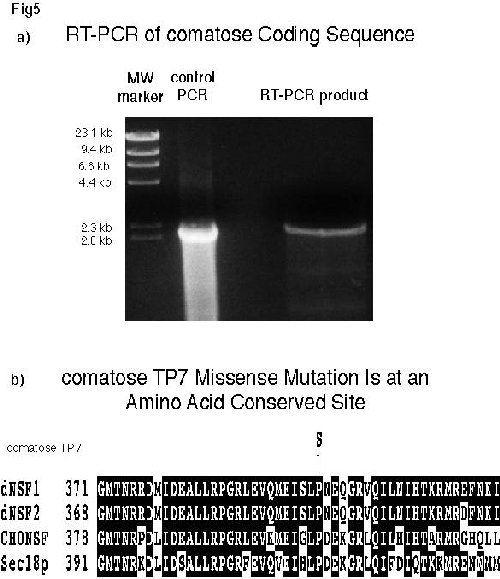 Fig. 5a: RT-PCR of comatose Coding Sequence. For control PCR, plasmid DNA carrying dNSF-1 cDNA was used as the template. In RT-PCR, reverse-transcribed mRNA from comatose flies was used as the template. The products were electrophoresed on 1% agarose plus ethidium bromide and DNA was detected by ultraviolet light.Fig. 5b: comatose TP7 Missense Mutation Is at an Amino Acid Conserved Site Comparison of the sequences of four NSF homologs demonstrates that the mutation in TP7 is a serine substitution for the conserved amino acid proline 397.
Fig. 5a: RT-PCR of comatose Coding Sequence. For control PCR, plasmid DNA carrying dNSF-1 cDNA was used as the template. In RT-PCR, reverse-transcribed mRNA from comatose flies was used as the template. The products were electrophoresed on 1% agarose plus ethidium bromide and DNA was detected by ultraviolet light.Fig. 5b: comatose TP7 Missense Mutation Is at an Amino Acid Conserved Site Comparison of the sequences of four NSF homologs demonstrates that the mutation in TP7 is a serine substitution for the conserved amino acid proline 397.
| <= Materials & Methods | RESULTS | Discussion & Conclussions => |
| Discussion Board | Next Page | Your Symposium |