Cell Biology Poster Session
| INABIS '98 Home Page | Your Session | Symposia & Poster Sessions | Plenary Sessions | Exhibitors' Foyer | Personal Itinerary | New Search |
Introduction
Accompanied with tumorigenesis, various genes are observed
to be altered in its expression, as well as activation of oncogene or inhibition
of tumor-suppressor genes etc. These marked differences in gene expression
between the normal cell and tumor are assumed to result in multiple phenotypic
change of tumor in morphology, metabolisms, and many often characters.
Therefore, elucidation of gene regulation in tumor cells is noteworthy
to understand the molecular mechanism of cell transformation.
Catalase [EC1.1.1.6] is a radical scavenging enzyme protecting
cells from oxidative stress, together with the other enzymes such as superoxide
dismutase and glutathione peroxidase. In mammalian tissues, the highest
level of catalase gene expression is found in the liver, kidney and erythrocytes
and the lowest level in connective tissues (1). Furthermore, the catalase
gene expression is known to be markedly reduced in cultured hepatoma cell
lines. It is interesting to analyze the mechanism of the catalase gene
expression.
Previously, we reported the G-rich silencer element located
-3.5kb upstream of the rat catalase gene (2). Furthermore, we also found
another negative regulatory region located in, -2799 to ?2652 bp, downstream
from the G-rich silencer element. In this study, we analyzed this region
-2799 to -2652 bp by EMSA and CAT assay, to disclose the fine mechanism
of negative regulation of the catalase gene in hepatoma cells.
Materials and Methods
I, Cell lines
AH66 cell is malignant and dedifferentiated hepatoma cell
line derived from an ascites rat hepatoma. Reuber cell is the rat minimal
deviation hepatoma cell line, and 3Y1 cell is an immortalized cell line
from Fischer rat embryonic fibroblast cell. All of these cell lines were
maintained in Dulbecco modified Eagle's medium supplemented with 10% fetal
calf serum.
II, Electrophoresis mobility shift assay (EMSA)
Preparation of nuclear extract from AH66 and Reuber cell was
performed according to Dignam's method (3). Probe DNA was end-labeled with
[ganma-32P]ATP and T4 polynucleotide kinase. EMSAs were performed as described
previously (4). DNA-protein complexes were resolved in 5% acrylamide:bis-acrylamide-TGE
gel containing 2.5% glycerol, and visualized by autoradiogrphy.
III, CAT(Chloramphenicol acetyltransferase) assay
Plasmid bearing truncated the 5'-flanking region of rat catalase
gene were prepared with restriction enzymes. CAT plasmid was constructed
by insertion of the truncated segments into the polylinker sites of pUC0CAT
and pCCAT-P11. pCCAT-P11 was constructed by insertion of the rat catalase
gene promoter region (-126 to -26 bp) into the polylinker sites of pUC0CAT.10ug
of these CAT constructs were transfected to 3Y1 by the calcium phosphate
precipitation procedure (5, 6), and the CAT activities were determined
with HPLC as described (7). Each transfection experiment was carried out
in triplicate, and the values of CAT activities, which deviated within
5%, were considered valid and are reported here.
Results
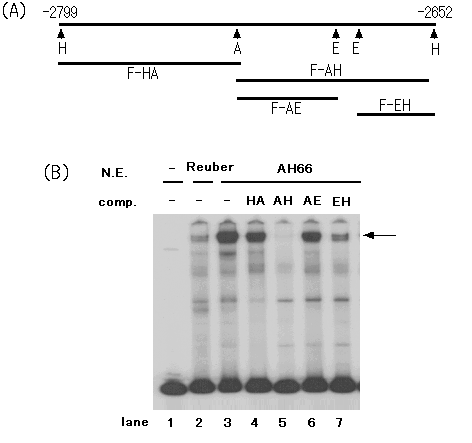
(1) Locating of binding protein in negative regulatory region
To study on negative regulatory element, located on -2799
to -2652 bp and designated F fragment, we determined protein binding
to this region by EMSA probed F fragment using nuclear extracts from hepatoma
cell lines, Reuber and AH66. The Reuber cell maintains catalase gene expression
at significant level as same as normal liver, while the AH66 cell extremely
deduced the catalase (2). As a result, a specific shift band was observed
in DNA complex with AH66 nuclear extract (Fig.1-B; lanes 2 and 3). Furthermore,
the truncated DNA fragments was determined by nuclear protein binding competition
assay (Fig.1-A). The DNA protein complex was strongly competed with F-AH
fragment (lane 4). Furthermore, F-EH fragment derived from F-AH fragment
also inhibited probe-protein complex formation in some extent. However,
competition by F-EH fragment was weak compared with F-AH fragment (lanes
5 and 7). Likewise, gel shift of F fragment was competed by F-HA and F-AE
fragment at quite low level (lanes 4 and 6). These results suggested that
nuclear protein in AH66 hepatoma cells composed DNA-protein complex with
F-EH fragment region, but scarcely with F-HA and F-AE fragments.
(2) Transcriptional activity of deletion mutant mutant
Next, to test the transcriptional activity of negative regulatory
elements in this region, CAT plasmids carrying various length of negative
element were constructed with F fragment and it derivatives. CAT activity
of pCCAT-F-EH showed strong repression in the rat catalase promoter. Further,
transcriptional activities by pCCAT-F-HA and pCCAT-F-AE were weakly repressed
compared with pCCAT-P11. These results suggested that a negative regulatory
element was located in AH fragment, paticulary in EH fragment.
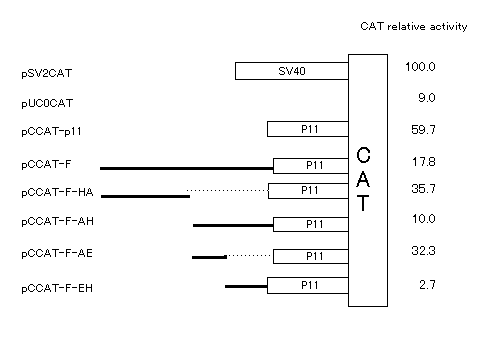 |
Fig. 2. CAT assay of various region in F fragment.
|
Discussion and Conclusion
Catalase activity is markedly reduced cultured hepatoma cell lines
(8, 9, and 10). Analysis of this phenomenon is assumed to disclose the
mechanism of carcinogenesis or dediffereaion of liver cell. From the results
of EMSA and CAT assay, we suggested that F-EH binding protein seemed a
transcriptional repressor of the rat catalase gene expression in rat hepatoma
cell lines.
Recent studies have identified a large number of proteins capable
of repressing transcription, and supporting evidence for this mechanisms
has been presented (11, 12). Transcriptional repression were classified
only two distinct functional mechanisms; position-independent silencing
and position-dependent one (13). Transcriptional silencer in the catalase
gene was necessary to elucidate the function in hepatocarcinogenesis.
References
1: Schoenbaum, G. R., et al. (1976) Catalase in The Enzymes 363-408.
Academic Press, New York.
2: Sato, K., et al. (1992) Mol. Cell. Biol. 12, 2525-2533.
3: Tamura, T., et al. (1989) Mol. Cell. Biol. 9, 3122-3126.
4: Fridell, Y., et al. (1992) Mol. Cell. Biol. 12, 4571-4577.
5: Chen, C., et al. (1987) Mol. Cell. Biol. 9, 1415-1425.
6: Gorman, C., et al. (1982) Mol. Cell. Biol. 2, 1044-1051.
7: Sato, K., et al. (1986) Mol. Cell. Biol. 6, 1032-1043.
8: Bozzi, A., et al. (1976) Mol. Cell. Biochem. 10, 11-16.
9: Marklund, S.L., et al. (1982) Cancer Res. 42, 1955-1961.
10: Potter, V.R. (1977) Biochemistry of cancer in Cancer Medicine 178-190, Lea and Febiger, Philadelphia.
11: Hanna-Rose., et al. (1996) Trends Genet. 12, 229-234.
12: Johnson, A. D. (1995) Cell. 81, 655-658.
13: Steven, O., et al. (1998) Biochem, J. 331, 1-14
| Discussion Board | Previous Page | Your Poster Session |