Cell Biology Poster Session
| INABIS '98 Home Page | Your Session | Symposia & Poster Sessions | Plenary Sessions | Exhibitors' Foyer | Personal Itinerary | New Search |
Introduction
The Couch spadefoot toad, Scaphiopus couchii, inhabits arid regions of the American southwest and spends as much as 9-10 months of each year burrowed underground in a state of dormancy, termed estivation. Long term survival is ensured by strong metabolic rate depression with oxygen consumption reduced to only 20-30 % of the resting rate when aroused and by adaptations that minimize the loss of body water as the soil dries out (1-3). Whereas considerable attention has been given to energy metabolism and water balance during amphibian estivation, little is yet known about the role of gene expression in promoting survival. Presumably, specific up- or down- regulation of selected genes is needed to optimize numerous metabolic processes to sustain long term dormancy. In the present study we differentially screened a cDNA library made from liver of 2 month estivated female toads in order to identify genes that were up-regulated during estivation and found a strong up-regulation of the gene for riboflavin binding protein.
Materials and Methods
All chemicals were of molecular biology grade and were obtained from Sigma Chemical Co., St. Louis, MO or New England Biolabs, Beverly, MA. Spadefoot toads were captured several days after emergence from estivation in July near Tucson, Arizona and were air-freighted to Carleton University where they were held at 15�C. Half of the toads were sacrificed within 24 h as controls; toads were killed by pithing and tissues were rapidly dissected, frozen in liquid nitrogen, and stored at -70�C. Remaining toads were placed in basins containing damp soil in an incubator at 15�C; toads burrowed into the soil and were allowed to estivate for 2 months. At the end of this time, toads were sacrificed; they had lost 29.4 % of their total body mass (equal to 35.7 % of total body water if all mass lost was water). Samples of frozen liver from female estivated toads were shipped on dry ice to Clontech (Palo Alto, CA) for the synthesis of a lambda Zap II cDNA library.
For cDNA probe synthesis, total RNA was extracted from liver of control and estivating female toads using Trizol reagent and then poly-A+ messenger RNA was purified using oligo-dT cellulose. 32P-Labeled, single-stranded cDNA probes were synthesized with random primers. For cDNA library screening a 1:125,000 dilution of the lambda library phage was incubated with 200 uL of an overnight culture of XL1-Blue MRF' host cells and then plated and grown overnight at 37�C. Two replicate lifts were made from of each plate using HYBond-N membrane and then lifts were probed with control versus estivated cDNA probes. Plaques showing greater signals with the estivation probe were cored, grown overnight, and re-screened to confirm differential expression of the clone and to obtain a pure phage stock. The Ex-Assist helper phage system was used to isolate potential differentially expressed inserts within pBluescript phagemids. The cDNA inserts were recovered from the plasmid by EcoR1/Xho1 digestion and electrophoresis in a 0.9% TAE/agarose gel. Potential positive inserts were purified from the agarose gel using Geneclean III kits. Random primer labeling was used to generate [gamma-32]P-dCTP-labeled probes for Northern blotting. Total RNA was extracted from samples of brain, gut, heart, lung, liver, kidney and muscle from both control and estivating female toads using Trizol reagent and then aliquots of each RNA extract were separated on agarose gels and hybridized using the radiolabeled probes. DNA sequencing was done in two ways: using Sequenase 2.0 (US Biochemicals) and by automated sequencing at the Core Facility for Protein/DNA Chemistry in the Biochemistry Department at Queen's University, Kingston, Ont. Sequence homologies were obtained from Genbank using the Blast program and additional analyses were performed using Lasergene sequence analysis software (DNASTAR).
Results
Of the clones isolated during primary and secondary screening of the cDNA library from liver of estivating female toads, one was confirmed as differentially expressed by Northern blotting. This clone had a 1.0 kb insert which annealed to a 0.7 kb band on a Northern blot. Expression of the mRNA transcript was 60 % higher in liver from estivated, compared with control, toads.
Sequencing revealed a full-length cDNA transcript of 1053 nucleotides (Figure 1). The deduced amino acid sequence of the longest open reading frame (708 nucleotides beginning at nucleotide residue 58) is also shown. This potentially encoded a protein of 235 amino acids with a predicted molecular weight of 26.6 kD. Homology search in Genbank indicated high similarity to riboflavin binding protein (RfBP).
 Fig. 1: Nucleotide sequence of the tRfBP cDNA clone and the deduced amino acid sequence (shown above the corresponding nucleotides) of the longest open reading frame. Nucleotide and amino acid (in bold) residues are numbered on the right and polyadenylation signal is underlined.
Fig. 1: Nucleotide sequence of the tRfBP cDNA clone and the deduced amino acid sequence (shown above the corresponding nucleotides) of the longest open reading frame. Nucleotide and amino acid (in bold) residues are numbered on the right and polyadenylation signal is underlined.
Figure 2 compares the amino acid sequence of the toad protein with RfBP from chicken oviduct (4) and turtle liver (5). Toad liver RfBP shared 50.4 % identity with the chicken protein and 49.6 % homology with the turtle protein whereas chicken and turtle proteins shared 70.2 % sequence homology.
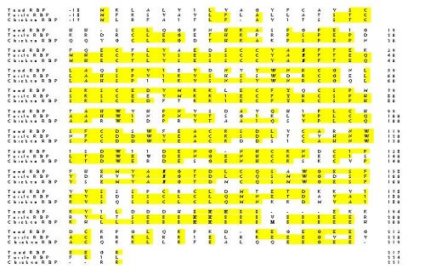 Fig. 2: Comparison of the deduced amino acid sequences of RfBP from spadefoot toad liver (Scaphiopus couchii), Chinese softshell turtle liver (Pelodiscus sinensis; 5), and domestic chicken oviduct (Gallus gallus; 4); Genbank accession numbers are AF102545, D49954 and J03922, respectively. Identical residues are boxed and conserved asparagine residues (glycosylation sites), tryptophan residues (involved in ligand binding), and phosphoserine cluster (recognition site for uptake by oocyte) are in bold. Amino acids in the mature protein are numbered on the right hand side and are preceded by the 17-18 residue signal sequence that is cleaved from the mature protein. A 4-residue insertion after the phosphoserine cluster was needed to align the C-terminal section of the toad protein with that of the other two species.
Fig. 2: Comparison of the deduced amino acid sequences of RfBP from spadefoot toad liver (Scaphiopus couchii), Chinese softshell turtle liver (Pelodiscus sinensis; 5), and domestic chicken oviduct (Gallus gallus; 4); Genbank accession numbers are AF102545, D49954 and J03922, respectively. Identical residues are boxed and conserved asparagine residues (glycosylation sites), tryptophan residues (involved in ligand binding), and phosphoserine cluster (recognition site for uptake by oocyte) are in bold. Amino acids in the mature protein are numbered on the right hand side and are preceded by the 17-18 residue signal sequence that is cleaved from the mature protein. A 4-residue insertion after the phosphoserine cluster was needed to align the C-terminal section of the toad protein with that of the other two species.
The areas of greatest variation between the three proteins were within the 17-18 residue signal sequence (which is cleaved from the mature protein) and in the final 25 residues near the C-terminus. In addition, a highly conserved region containing 8 phosphoserine residues, that is present in the turtle and chicken proteins, was reduced to only 3 potential serine phosphorylation sites in toad RfBP.
Northern blots using total RNA isolated from brain, gut, heart, kidney, and leg skeletal muscle of estivating and control toads and probed with the liver clone (termed tRfBP) failed to detect expression of this gene in four tissues. However, transcripts of tRfBP were detected in hind leg skeletal muscle from estivating toads but with transcript levels only about 10 % of the corresponding amount in liver of the same animals.
Discussion and Conclusion
1. Differential gene expression during estivation:
The isolation of only a single clone as differentially up-regulated during estivation was unexpected as we had predicted that numerous changes in gene expression would be needed to accomplish the biochemical adjustments needed for estivation. However, this result might be explained in one of two ways.
(a) The overall lower metabolic rate in the estivating state would mean that most genes are down-regulated compared with their status in control animals. Against this background, a functional "up-regulation" of selected genes that support estivation could be achieved by simply not changing their level of expression in the estivating, compared with control, states.
(b) Upregulation of those genes that facilitate the establishment of dormancy or the initiation of estivation-specific functions (e.g. urea synthesis) would likely be complete long before two months had passed such that toads were in steady-state dormancy by this time. Hence, identification of RfBP up-regulation suggests a very specific function for this protein during long term dormancy.
2. RfBP function:The deposition of water-soluble vitamins into eggs of birds and reptiles or into the mammalian fetus is facilitated by vitamin binding proteins. RfBP was first identified in bird eggs and is synthesized by liver (yolk protein) and oviduct (white protein). Reptiles show only the yolk form (5). Our study is the first to find RfBP in an amphibian and hence is interesting in both functional and evolutionary ways.
Elevated RfBP mRNA (60 % higher than in controls) in liver of 2 month estivated toads suggests long term changes in the vitamin metabolism of toads. Most likely, increased RfBP mRNA in liver implies increased RfBP protein synthesis and secretion into plasma. There RfBP would bind riboflavin and load it into maturing eggs of female toads. Spadefoot toads are explosive breeders; the first heavy rains of summer bring thousands of toads to the surface and breeding is completed within 24 h after emergence. Thus, eggs must be fully mature when toads emerge from estivation and egg development must be completed during dormancy.
The source for the riboflavin that is scavenged by RfBP might be vitamins that are released of a result of wasting of skeletal muscle mass during extended dormancy. Nearly 20 % of body protein reserves are lost during estivation (6). Thus, vitamins lost from muscle or other organs could be recycled into eggs. In this regard, the results of northern blotting that showed the presence of RfBP mRNA in leg muscle of estivated, but not control, toads are interesting.
3. Toad RfBP stucture:RfBP from bird and reptile sources is a monomeric phosphoglycoprotein of about 30 kD, similar to the 26.6 kD suggested for toad RfBP.
Figure 2 shows that toad liver RfBP is very similar to the chicken and turtle proteins. All three have open reading frames that code for 235-242 amino acid residues and include a 17-18 amino acid signal sequence, starting with methionine and ending with cysteine, that is cleaved from the mature protein. The chicken cDNA also codes for two arginine residues at the C terminus that are not present in the mature protein of 219 amino acids but whether C terminal processing also occurs with toad or turtle RfBP is not known.
Amino acid residues known to be essential to RfBP function are all conserved in toad RfBP. These include 18 cysteine residues that form 9 disulfide bridges (7), 2 asparagine glycosylation sites (Asn35 and 145 in toad, Asn36 and 147 in chicken and turtle) and 6 tryptophan residues that are thought to be involved in ligand binding (8). The ligand binding domains runs from the N-terminus to about Cys169 in chicken RfBP (Cys167 in toad)(7).
A highly phosphorylated region extends from amino acids 186 to 197 in the chicken and turtle protein and contains eight phosphoserines interspersed with glutamate residues. However, the toad protein shows only three serines in this region, spaced by two glutamate residues. This is the only major structural difference between the toad protein and the bird/reptile proteins. RfBP transport across the yolk membrane in bird eggs is facilitated by a specific carrier that recognizes the region of eight phosphoserines (9). Hence, the much smaller sequence of serine residues in the toad protein, suggests that the recognition and binding of RfBP to toad oocytes may be quite different in amphibians.
4. Conclusions:This study presents the first use of cDNA library screening technology to investigate the changes in gene and protein expression that facilitate amphibian estivation. Our findings show the enhanced expression of the gene for riboflavin binding protein in female spadefoot toads after two months of dormancy. The data provide the first confirmation of the presence of RfBP in an amphibian species. The probable role of the protein is in vitamin loading into eggs so that these are fully mature in anticipation of the explosive breeding and egg-laying that occurs immediately when toads are aroused from dormancy.
References
1. Storey, K.B., and J.M. Storey (1990) Metabolic rate depression and biochemical adaptation in anaerobiosis, hibernation and estivation. Quart. Rev. Biol. 65: 145-174.
2. McClanahan, L. (1967) Adaptation of the spadefoot toad, Scaphiopus couchii, to desert environments. Comp. Biochem. Physiol., 20:73-99.
3. Seymour, R.S. (1973) Energy metabolism of dormant spadefoot toads (Scaphiopus). Copeia 1973:435-445.
4. Zheng, D.B., H.M. Lim, J.J. Penes, and H.B. White (1988) Chicken riboflavin-binding protein: cDNA sequence and homology with milk folate-binding protein. J. Biol. Chem., 263:11126-11129.
5. Hamajima, S., and S. Ono (1995) Sequence of a cDNA encoding turtle riboflavin-binding protein: a comparison with avian riboflavin-binding protein. Gene, 164:279-282.
6. Jones, R.M. (1980) Metabolic consequences of accelerated urea synthesis during seasonal dormancy of spadefoot toads, Scaphiopus couchi and Scaphiopus multiplicatus. J. Exp. Zool., 212:255-267.
7. Monaco, H.L. (1997) Crystal structure of chicken riboflavin-binding protein. EMBO J., 16:1475-1483.
8. Blankenhorn, G. (1978) Riboflavin binding in egg white flavoprotein: the role of tryptophan and tyrosine. Eur. J. Biochem. 82:155-160.
9. Sooryanarayana, S.S., P.R. Adiga, and S.S. Visweswariah (1998) Identification and characterization of receptors for riboflavin carrier protein in the chicken oocyte. Role of the phosphopeptide in mediating receptor interaction. Biochim. Biophys. Acta, 1382:230-242.
| Discussion Board | Previous Page | Your Poster Session |